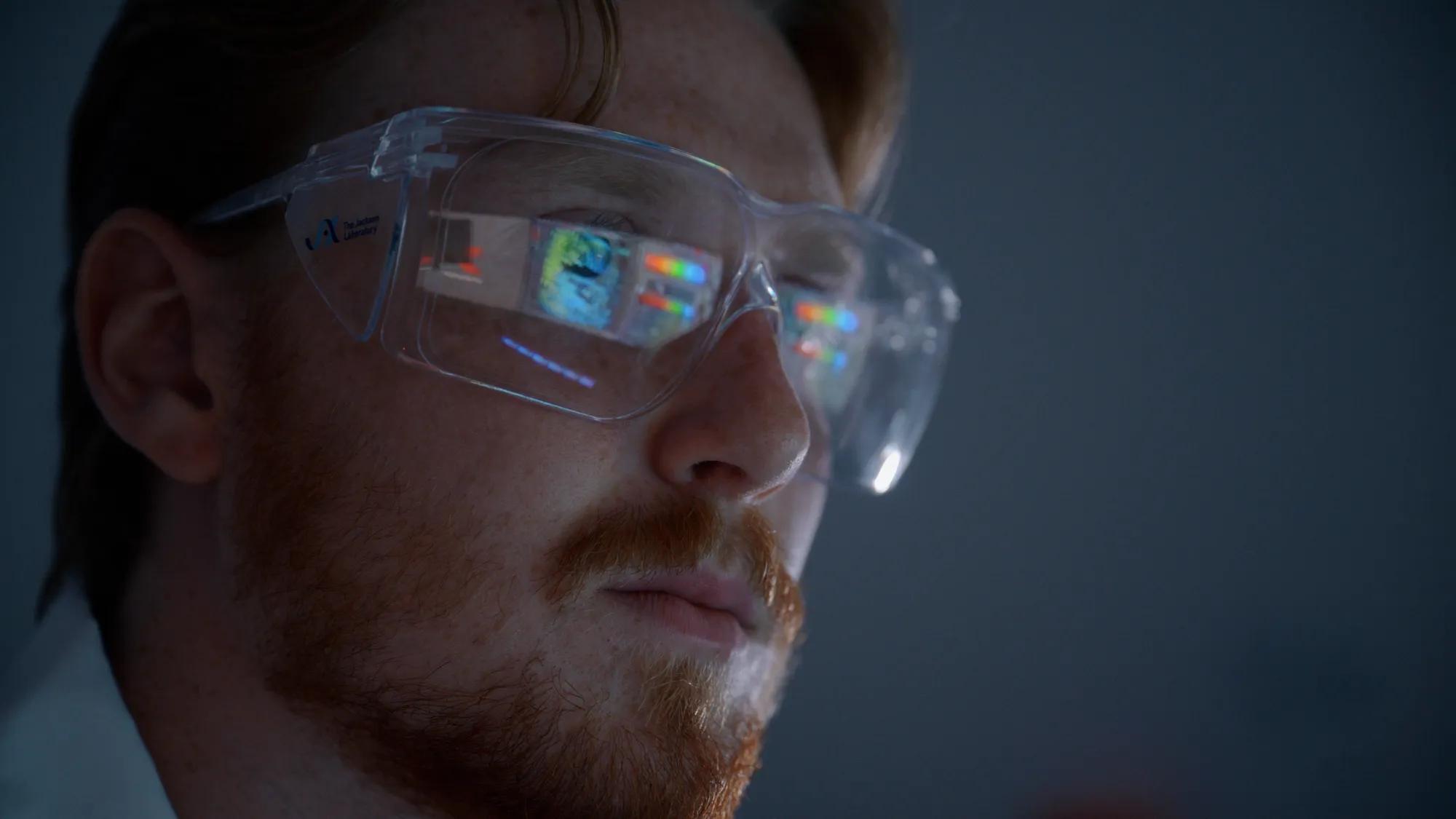
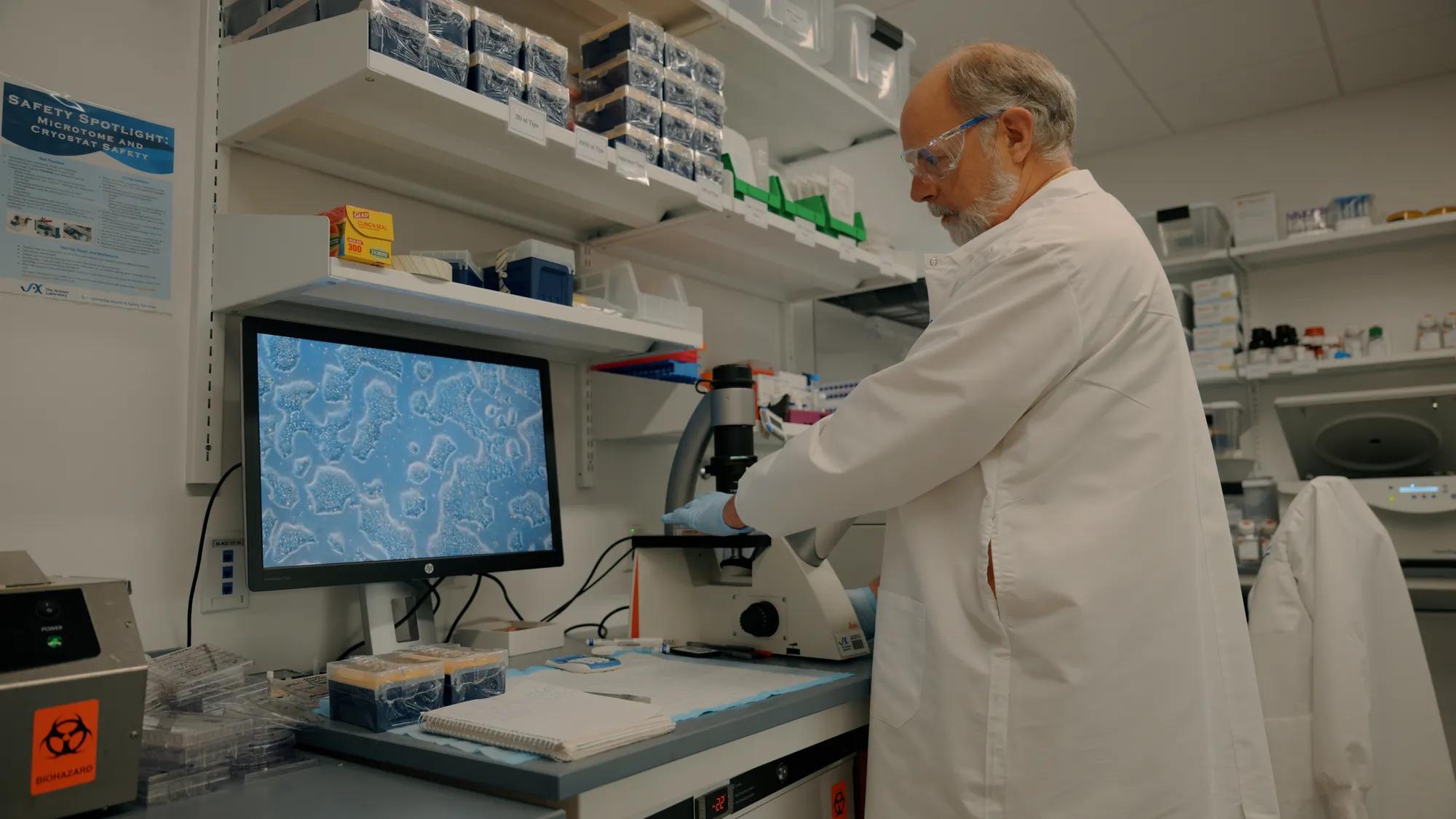
More than two decades after the human genome was first sequenced, scientists still don't know what many of its approximately 20,000 protein-coding genes actually do. While a handful of genes have been intensely studied, most remain biological mysteries, leaving major gaps in the understanding of how human biology works and how it goes wrong in disease.
That’s the challenge the MorPhiC consortium aims to solve. Backed by the National Human Genome Research Institute, MorPhiC (short for Molecular Phenotypes of Null Alleles in Cells) is a nationwide initiative to determine the function of every human gene by “turning off” each one and seeing how living cells respond.
Helping lead this effort is a team at The Jackson Laboratory (JAX), drawing on decades of experience from its role in the Knockout Mouse Project. Now, JAX researchers are applying those lessons to human cells—generating precise gene knockouts, developing scalable methods to study the effects of these changes, and sharing the resulting cell lines as an open resource with the rest of the global scientific community.

To do this with the entire genome is a very complicated and ambitious project. But this is basic information that we need to know to understand human biology.”
- Bill Skarnes
From mouse to human
JAX has long been at the forefront of efforts to understand what genes do by systematically studying their functions. JAX researchers including Skarnes have played major roles in the International Mouse Phenotyping Consortium, which has studied the effect of thousands of gene knockouts on mice. That experience positioned the lab to take on an equally ambitious task with MorPhiC: defining the function of human genes, not in animal models, but directly in human cell systems.
Unlike mice, human cells require sophisticated lab-based systems to mimic development. Creating the new gene knockouts starts by disrupting the genes in induced pluripotent stem cells (iPSCs), which have the potential to yield many other human cell types. Then, the cells are coaxed to differentiate into adult cells of interest, like brain or heart cells.
“In the mouse, the organism naturally develops all its tissues,” said JAX Professor Paul Robson. “But in human cell models, we have to manipulate media composition and differentiation strategies to derive cell types of interest. That’s why the recent advances in comprehensive iPSC differentiation strategies have been critical to making this kind of large-scale project feasible.”
New gene editing technologies are also a boon to the project. Across MorPhiC, different institutions are turning to different strategies for disrupting genes—cutting the DNA so cells can no longer read the gene at all, blocking the RNA that is required to translate a gene into a protein, or degrading the corresponding protein once it has been made.
The JAX group primarily uses CRISPR to edit the gene itself, and has developed a precise, reversible method of disruption. This allows researchers to “rescue” the gene—restoring its function—to confirm that any observed changes in the cell were truly caused by the knockout.
“This kind of reversibility is essential,” said Skarnes. “It’s the best way to confirm that the phenotype you’re seeing is caused by the gene you targeted, and not by some unintended effect of the editing process.”
250 genes and counting
MorPhiC’s first phase focuses on 1,000 genes, with JAX responsible for 250 of them. The genes were selected for both their biological interest and their uniquely human qualities—such as genes involved in early human brain development or the formation of the placenta.
So far, the JAX team has successfully engineered cells lacking 170 of the genes and is systematically analyzing the effects of each one—a process that the consortium outlined in a February paper in Nature. At the same time, the group publicly released data on the first 71 gene knockouts across multiple cell types.
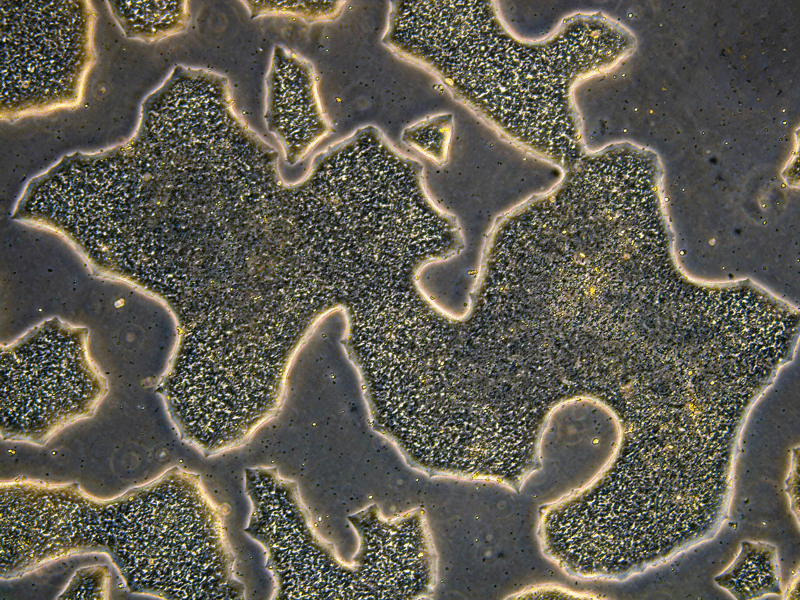
The efforts to characterize each cell line go well beyond checking whether cells live or die. JAX scientists are using RNA sequencing and other molecular assays to profile how a knockout changes gene expression and cell identity. Some early results have already uncovered intriguing biology.
For instance, researchers studying placental cells found that several genes involved in oxygen sensing also play key roles in how cells develop and differentiate—findings that may help explain pregnancy complications like preeclampsia. Another set of genes turned out to be essential for steroid hormone production in the placenta, a function not shared with mice.

These are genes that have evolved recently in humans and are linked to adult diseases like cancer and gout. It’s a perfect example of how understanding basic developmental biology can shed light on health and disease across the lifespan.”
- Paul Robson
A lasting resource for the scientific community
One of MorPhiC’s most important contributions won’t be the data alone—it will be the cell lines themselves. Each engineered cell line is banked and shared through JAX, enabling other scientists to study the same gene knockouts in different contexts or cell types.
“We expect phenotyping to go on for years,” said Skarnes. “New methods will come online, new disease models will emerge. These cell lines are a permanent resource to support those future discoveries.”
The project is also driving innovation on the computational front. A data analysis team led by JAX’s Brian White, a senior computational scientist, is evaluating how entire gene networks respond to the knockouts. This work could help map gene interactions and pathways, even for genes not directly perturbed.
“If two genes respond similarly to a wide range of knockouts, we can infer that they’re functionally related—maybe part of the same pathway,” said White. “That kind of insight can be incredibly useful for understanding disease mechanisms and identifying therapeutic targets.”
By revealing what genes and gene networks do in normal development, the project helps scientists interpret the effects of mutations found in patients, accelerating research into rare diseases, cancer, and more, he added.
What’s next
Ultimately, the consortium aims to expand from 1,000 genes to the more than 20,000 protein-coding genes in the human genome. Whether that will happen depends in part on the success of this initial phase—and on the continued support of funding agencies and the scientific community.
For now, the JAX team remains focused on building a high-quality, scalable platform for human gene discovery—one knockout at a time. They are also expanding the cell lines being used to create each gene knockout; rather than relying only on cells from a Northern European male, they’ll also begin using cells from male and female donors of African ancestry to assess how genetic background influences gene function.
Learn more

The Skarnes Lab
Expert genetic research and mouse models at JAX drive scientific breakthroughs, ensuring precision and reliability in advancing medical discoveries.
View more
JAX in Motion
JAX in Motion seeks to inspire and move audiences through a series of short documentaries. Presenting the groundbreaking research and discoveries being made by the exceptional scientific researchers at The Jackson Laboratory.
View more